Ximmer User Guide
Introduction
Ximmer is a tool designed to help users of targeted high throughput (or “next generation”) genomic sequencing data (such as exome data) to accurately detect copy number variants (CNVs). Ximmer is not a copy number detection tool itself. Rather, it is a framework for running and evaluating other copy number detection tools. It offers three essential features that users of CNV detection tools need:
- An analysis pipeline that automatically configures and runs a variety of well known CNV detection tools
- A simulation tool that can create artificial CNVs in sequencing data for the purpose of evaluating performance
- A visualisation and curation tool that can combine results from multiple CNV detection tools and allow the user to inspect them, along with relevant annotations.
We created Ximmer because although there are very many CNV detection tools, they can be hard to run and their performance can be highly variable and hard to estimate. This is why Ximmer builds in simulation: to allow a quick and easy estimation of the performance of any tool on any data set.
Want see how Ximmer results look without running it? Try out the live example report.
A picture of Ximmer’s accuracy report is below:
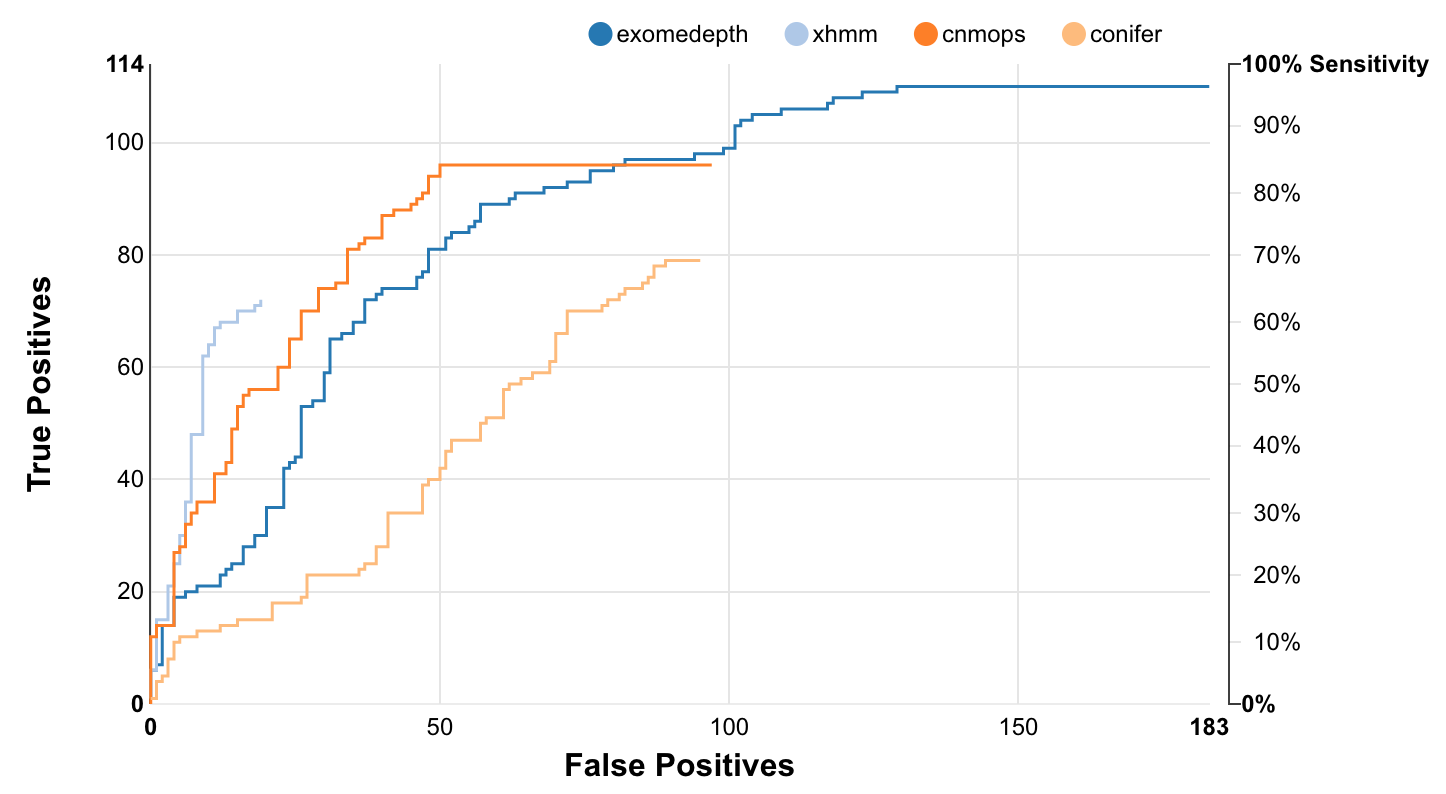
Installation
Ximmer can be installed both natively and using a docker container. Refer to the Install Guide for detailed instructions about how to get Ximmer running.
One Time Configuration
The default installation works as-is if you are running Ximmer in a stand alone environment. If you want to run on a computational cluster or configure other aspects in more detail (for example, the exact locations of tools such as R, or python that are used, the level of concurrency to employ, or to set up Ximmer to work on a computational cluster), see the Configuration documentation for more details.
Running Ximmer
See information about how to run Ximmer in the Running section.